
Last week I started this two-part blog series about increasing the speed of Python through Cython and PyPy. While we already discussed the effects of Cython, I wanted to continue the discussion with a look at PyPy and then a contrast of both PyPy and Cython.
Contents
What is PyPy and how fast is it?
PyPy is an alternative Python interpreter and just-in-time (JIT) compiler that is highly compatible (subject to a few caveats) with the CPython interpreter. It is designed for speed and efficiency and uses a tracing JIT compiler to optimize frequently executed parts of the program at runtime, thus increasing the execution speed.
PyPy runs regular Python programs; in general, no modifications need to be made to Python code to allow it to run on PyPy. Owing to the nature of the JIT compiler, it needs time to analyze frequently executed parts of the program, so it works best on programs that run for longer than a few seconds.
Perhaps the biggest hurdle to using PyPy in a production system is its lack of support for CPython extension modules. According to the PyPy documentation, extension module support is experimental and often runs much slower than in CPython. This means that some third-party libraries that have C extension modules might not work on PyPy — the scientific computing library NumPy is a notable example. Having said this, PyPy is able to run the majority of popular Python frameworks and libraries, such as Django, Flask, and SQLAlchemy.
The PyPy Wikipedia page and the PyPy website provide plenty of information on what PyPy is and how to use it; the latter also has a nice section that shows a list of benchmarks and their speed improvements over CPython.
How Cython vs CPython Works
Test Program: Numerical Integration
My main objective in doing this exercise was to understand how the tools work, and to be able to compare the differences in syntax and code structure between them. With that in mind, I chose to write a simple numerical integration program, which is based on an example given in the Cython documentation. The full project source, including installation and Cython build instructions, can be found on Bitbucket.
In the following code listing, the pure Python implementation is on the left, while the Cython version is on the right.
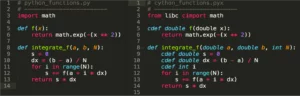
Cython vs Python Code
The Cython code differs from pure Python in the following ways:
Cython modules have a .pyx file extension instead of .py. The Cython build process translates them into intermediate C source files then compiles them using the system’s C compiler.
Cython code looks like Python code with added type declarations. Type declarations for variables, function arguments, and return values are optional, but adding them gives Cython enough information to translate into more optimized C code.
Cython allows easy linking to C libraries. At line 3, I’ve used the cimport statement to import the native libc math library instead of the Python math library, which improves performance.
Cython allows native C functions, which have less overhead than Python functions when they are called, and therefore execute faster. At line 5, I’ve defined f as a native function using the cdef statement. A downside to using native cdef functions is that they are not accessible by Python code outside the Cython module – effectively they are private.
Lines 9-11 in the integrate_f function define the variables. Defining i as an integer at line 11 allows Cython to translate the for loop at line 12 into a native C for loop, which improves performance. If the type of i was not specified, the loop would execute more slowly because the type of i would have to be checked at each iteration.
CPython, Cython & PyPy Speed Results
Benchmark Results
I tested the following three implementations of the algorithm:
- CPython: The pure Python implementation running in the standard CPython interpreter
- CPython + Cython: The Cython implementation running in the standard CPython interpreter
- PyPy: The pure Python implementation running in the PyPy interpreter
In order to measure the speed of program execution for each implementation, I wrote a program to call integrate_f 500 times using an N value of 50,000, and recorded the execution time over several runs.
I ran the tests on a 2015 Macbook Pro, using CPython 2.7.9, Cython 0.24, GCC 4.2.1 and PyPy 2.5.1 (compatible with CPython 2.7.9).
The following table shows the benchmark results:
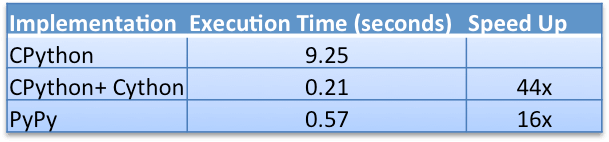
The CPython + Cython implementation is the fastest; it is 44 times faster than the CPython implementation. This is an impressive speed improvement, especially considering that the Cython code is very close to the original Python code in its design.
The PyPy implementation is 16 times faster than the CPython implementation and about 3 times slower than the Cython implementation. This is fascinating since PyPy is running the exact same pure Python code as the CPython implementation – it shows the power of PyPy’s JIT compiler.
Summary of Cython vs CPython vs PyPy
And The Winner Is…
I’ve enjoyed learning about Cython and PyPy and am impressed by the speed gains that are possible with only a small amount of programming effort. Speed is certainly addictive, and it’s hard to go back to running my 9-second Python program when I know the Cython one runs in 0.2 seconds!
My numerical integration test program is, of course, a toy problem, and it should be remembered that benchmarking results can vary considerably depending on the algorithms. In a real project, I would likely use NumPy or another specialized third-party library to perform my calculations.
CPython C extension modules, the Cython language and the PyPy interpreter are deep and interesting subjects in their own right and I’ve only scratched the surface in this blog post. Each tool has its own pros and cons, and may be a great fit for one project and a bad fit for another.
Having experimented with both tools, I am more likely to use Cython for speed-critical parts of programs in the future. The slow parts of the code can be identified by profiling and refactoring into a Cython module, which could be written to only contain the functions that needed to be optimized.
PyPy seems like it would be more of a quick fix, swapping out the CPython interpreter for PyPy without modifying a project’s existing code base. I would be concerned, however, that somewhere down the line a third-party library would be required for some functionality, and that it would turn out to be incompatible with PyPy. You could very possibly paint yourself into a corner.
I hope that I’ve managed to provide a valuable introduction to the world of Cython and PyPy. I already knew that Python was awesome — now I know that it can be fast too!
If you enjoyed this blog and are curious about our job openings, check out our careers page below.
